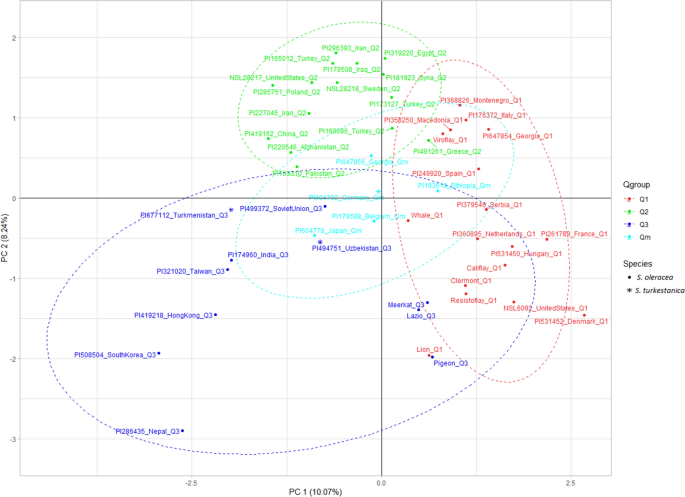
Genome-wide analysis of SSR and ILP markers in trees: diversity profiling, alternate distribution, and applications in duplication | Scientific Reports
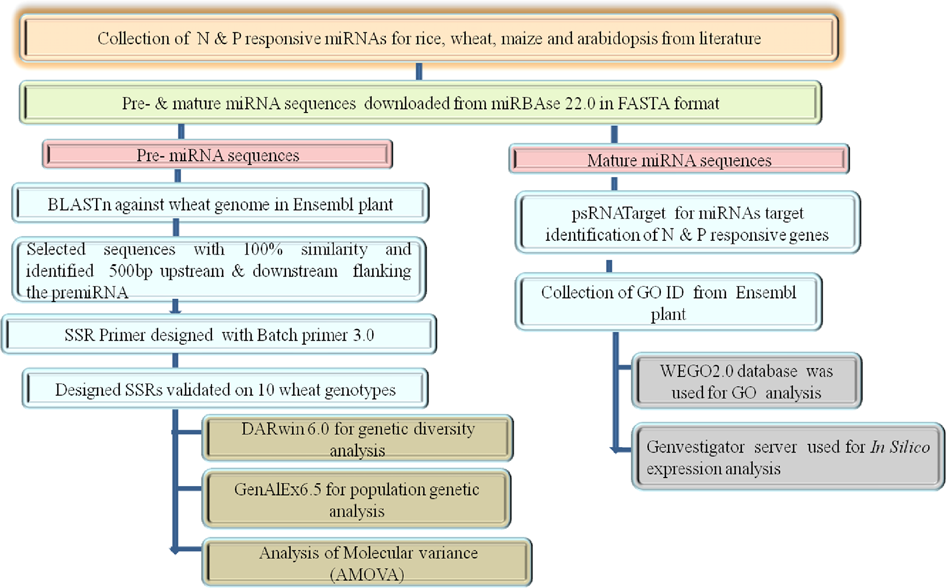
Development and characterization of nitrogen and phosphorus use efficiency responsive genic and miRNA derived SSR markers in wheat | Heredity

PDF) Development of Simple Sequence Repeat Markers in Hazelnut (Corylus avellana L.) by Next Generation Sequencing and Discrimination of Turkish Hazelnut Cultivars using SSR Barcodes | ResearchGate
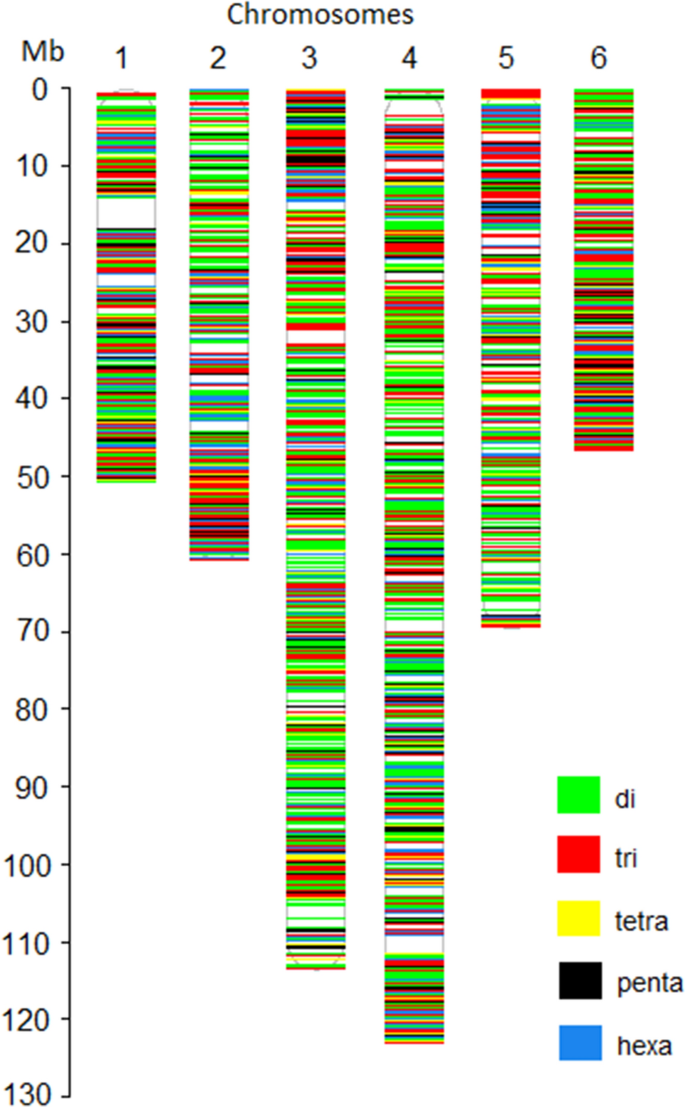
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

Phylogenetic tree analysis of gene based SSR marker data analysis using... | Download Scientific Diagram

Microsatellite markers are powerful tools for discriminating among olive cultivars and assigning them to geographically defined populations
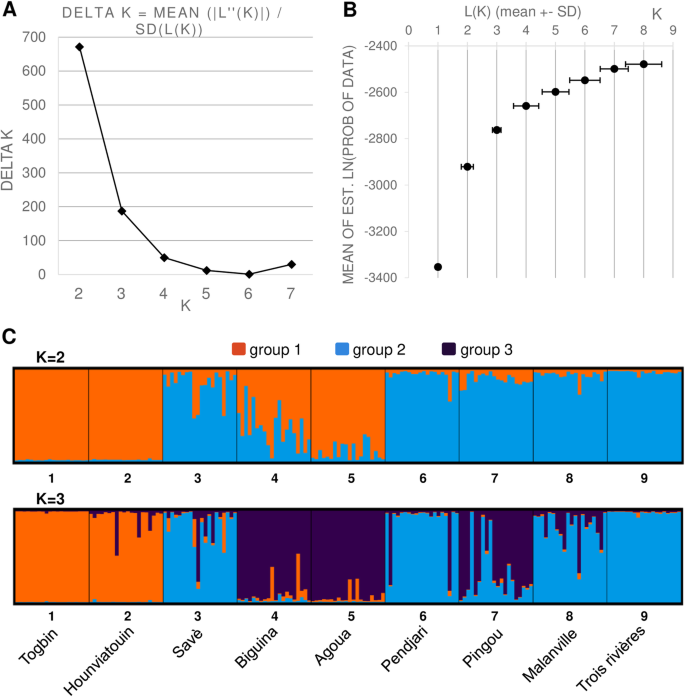
Transferability, development of simple sequence repeat (SSR) markers and application to the analysis of genetic diversity and population structure of the African fan palm (Borassus aethiopum Mart.) in Benin | BMC Genomic

Development of Genome-wide SSR Markers for Physical Map Construction with PCR-based Polymorphic SSRs in Jute (Corchorus Spp.) | SpringerLink
Identification of a minimal microsatellite marker panel for the fingerprinting of peach and nectarine cultivars

PDF) Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions

PDF) SureSawitTM TRUE-TO-TYPE – A HIGH THROUGHPUT UNIVERSAL SINGLE NUCLEOTIDE POLYMORPHISM PANEL FOR DNA FINGERPRINTING, PURITY TESTING AND ORIGIN VERIFICATION IN OIL PALM
Development of 16 novel EST-SSR markers for species identification and cross-genus amplification in sambar, sika, and red deer | PLOS ONE

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

Microsatellite markers are powerful tools for discriminating among olive cultivars and assigning them to geographically defined populations
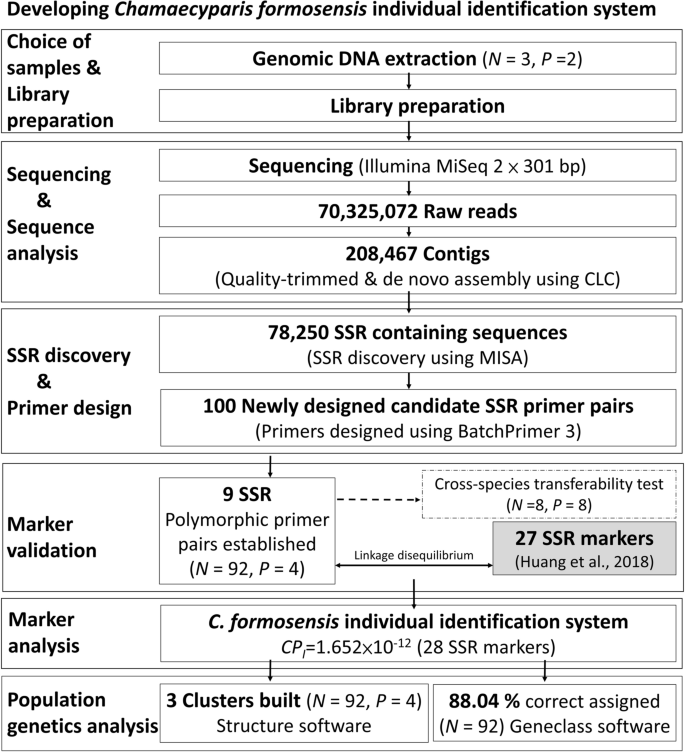
SSR individual identification system construction and population genetics analysis for Chamaecyparis formosensis | Scientific Reports

Transcriptome wide SSR discovery cross-taxa transferability and development of marker database for studying genetic diversity population structure of Lilium species | Scientific Reports

Development of Genome-wide SSR Markers for Physical Map Construction with PCR-based Polymorphic SSRs in Jute (Corchorus Spp.) | SpringerLink
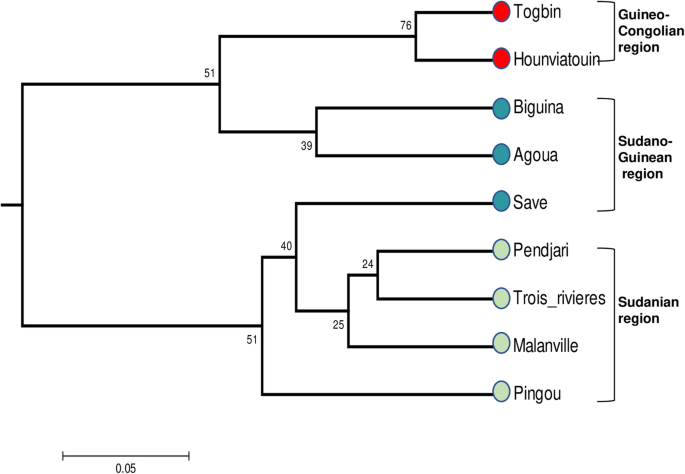
Transferability, development of simple sequence repeat (SSR) markers and application to the analysis of genetic diversity and population structure of the African fan palm (Borassus aethiopum Mart.) in Benin | BMC Genomic
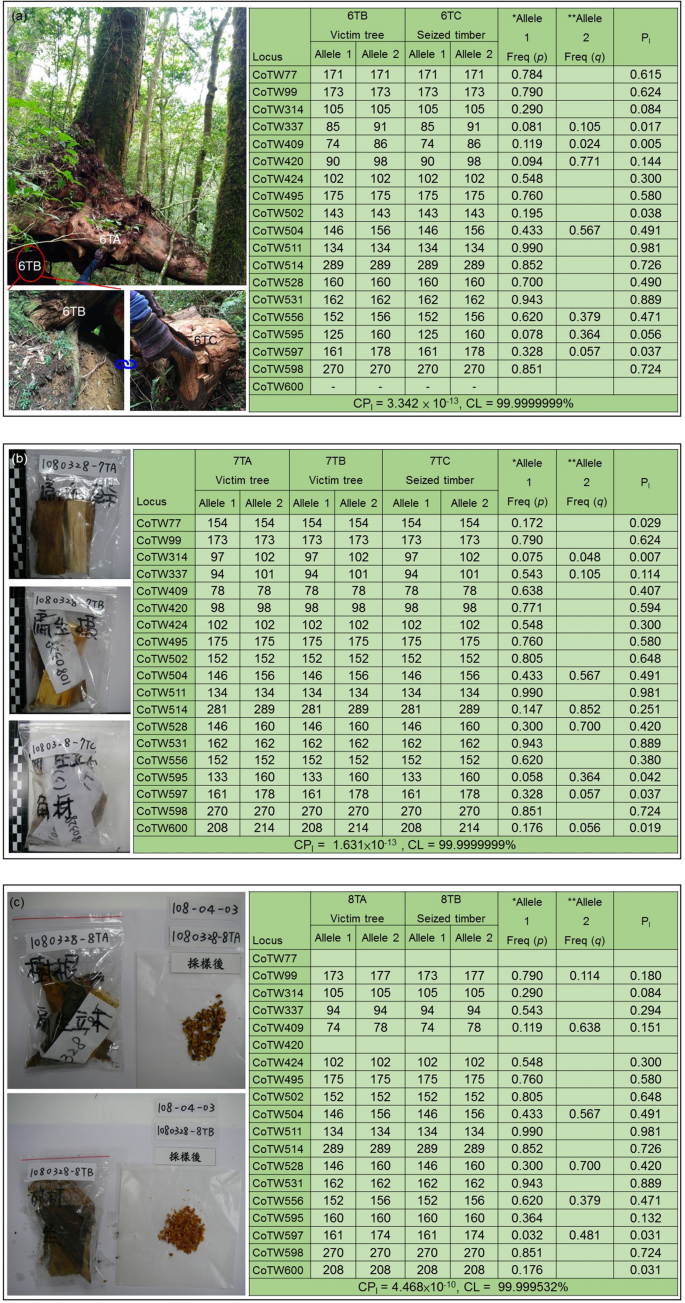
Development and technical application of SSR-based individual identification system for Chamaecyparis taiwanensis against illegal logging convictions | Scientific Reports
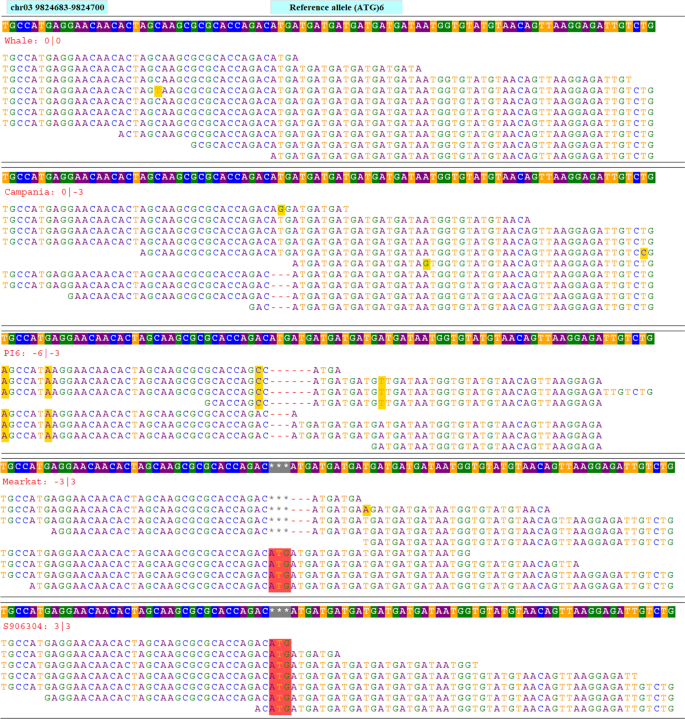
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

PDF) Identification of a minimal microsatellite marker panel for the fingerprinting of peach and nectarine cultivars
Criteria for evaluating molecular markers: Comprehensive quality metrics to improve marker-assisted selection | PLOS ONE
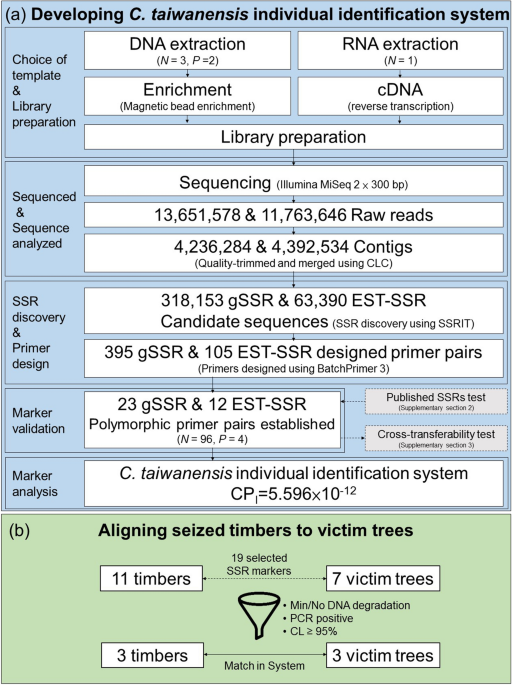
Development and technical application of SSR-based individual identification system for Chamaecyparis taiwanensis against illegal logging convictions | Scientific Reports

PDF) Informative SSR markers for commercial variety discrimination in watermelon (Citrullus lanatus)